The Decision Operating System for Drug Discovery.
With BioBox, biopharma teams don’t just manage data, they make scientific reasoning programmable.
















Why Drug Discovery Teams Choose BioBox
Your Data. Your Graph. Your Advantage.
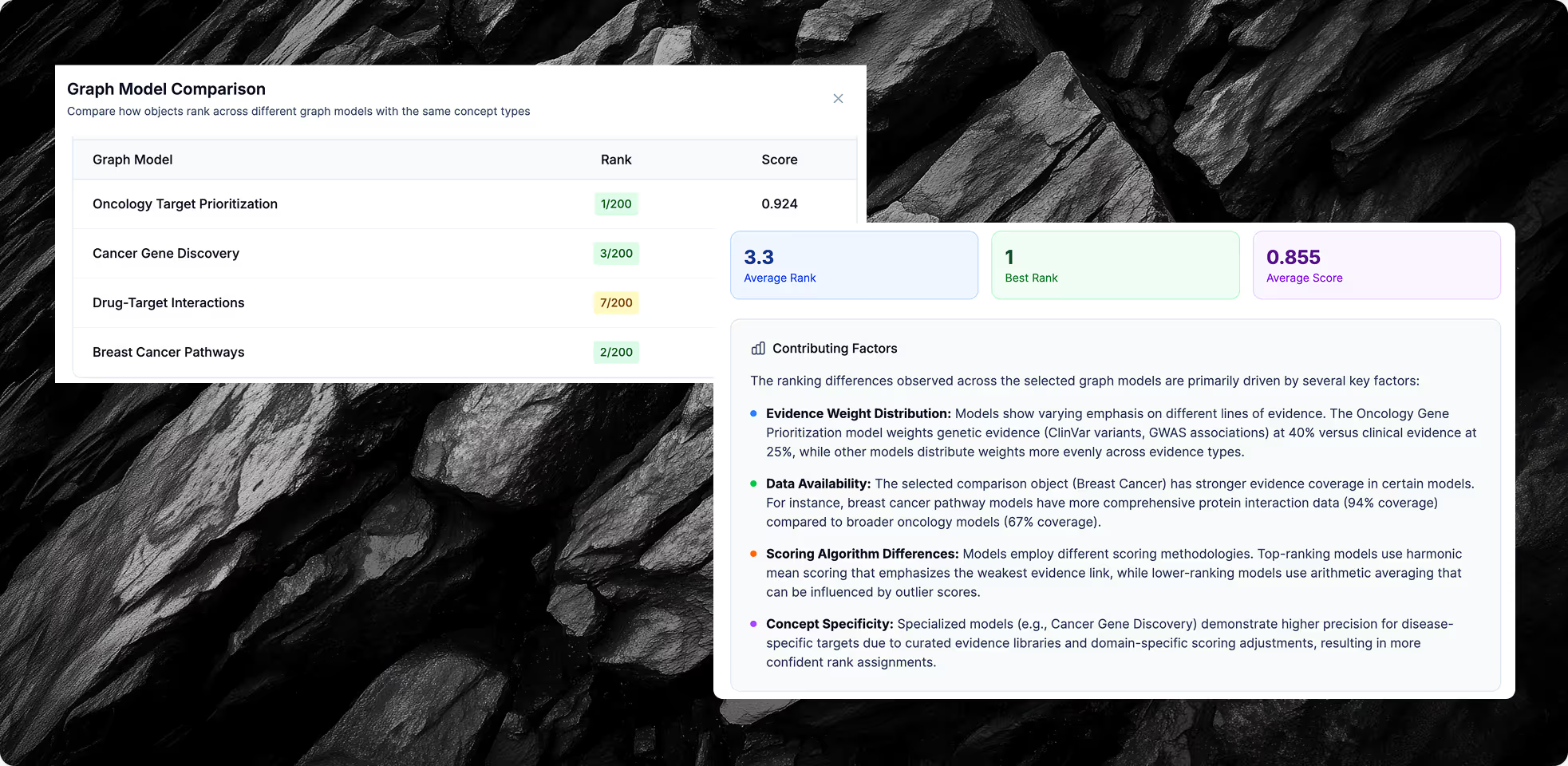
Reasoning at Scale
Powering Agentic Science
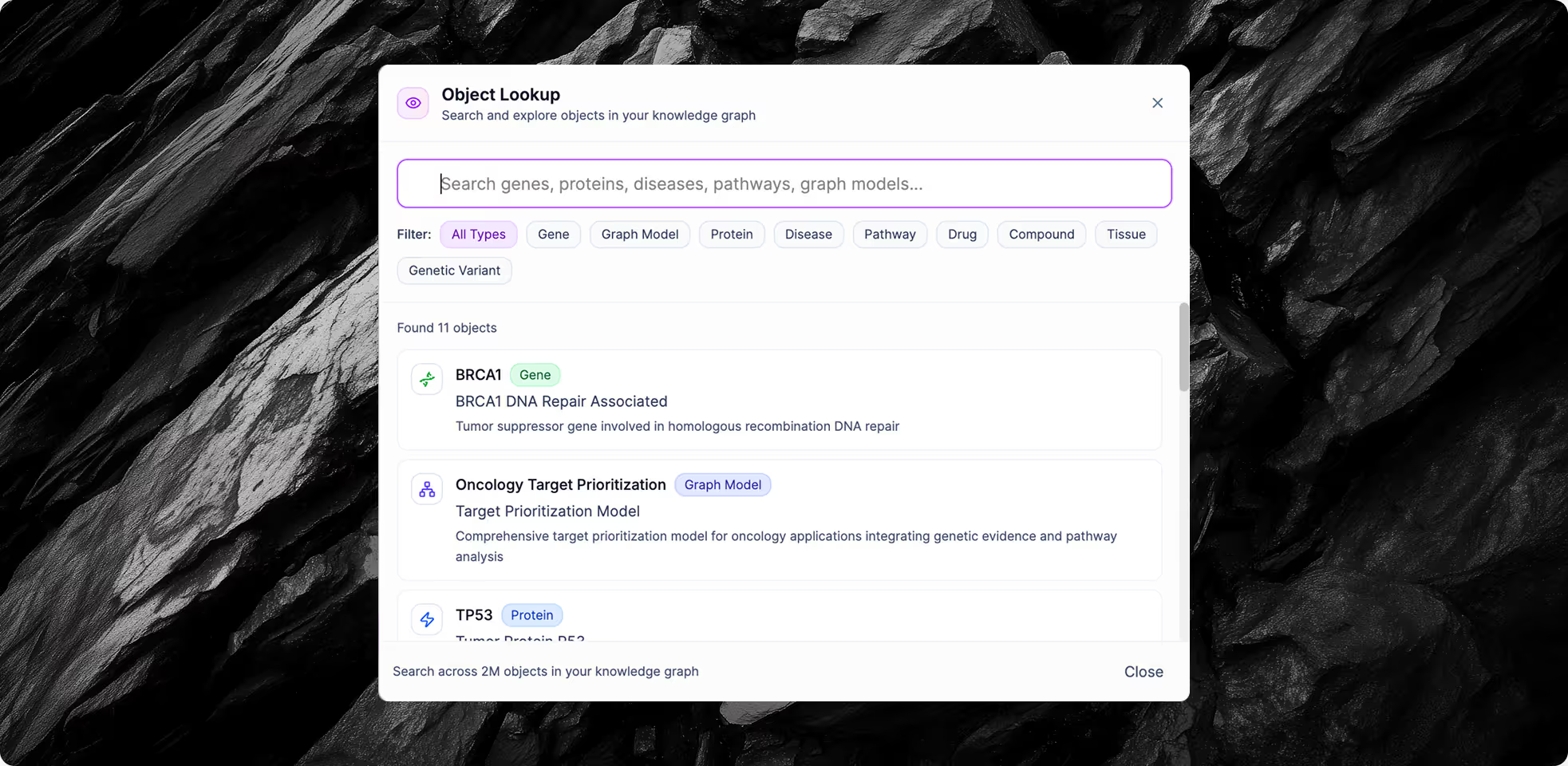
End-to-End Integration
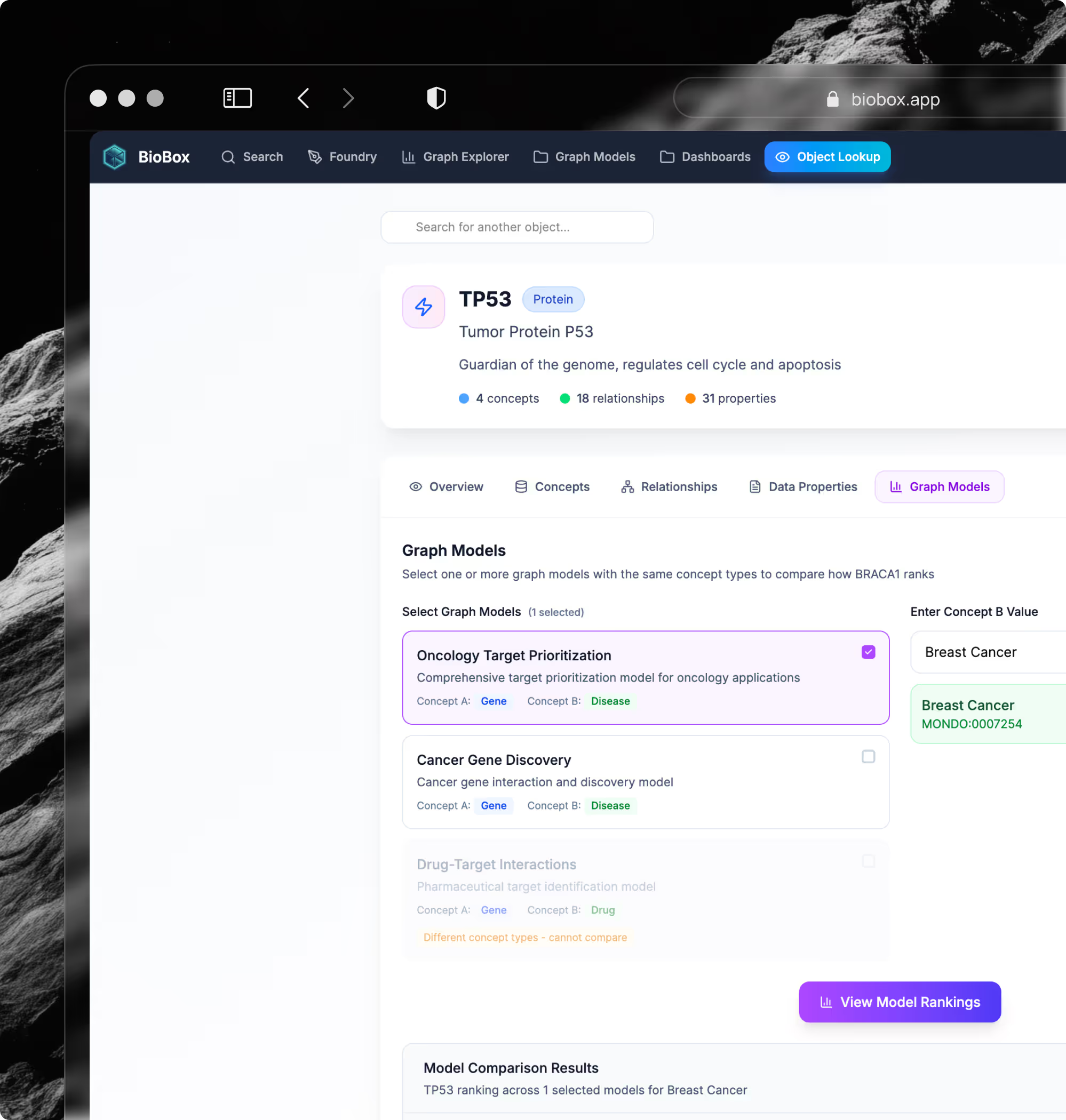
More Clarity. Less Friction. Better Results.
Real World Impact.
Everything You Need to Know—Upfront
Every graph built is fully custom, using your an ontology developed by you and built around your data. The graphs conform to your science. BioBox graphs are data-driven and constructed using real-world experimental observations, not blind literature mining.
Each engagement varies depending on client needs. Contact us to learn more.
You'll have a choice to purchase the software license. Even if you don't you get to keep your knowledge graph.
Yes, BioBox is SOC 2 Type II, ISO27001 and GDPR compliant and supports role-based access, audit logging, and data encryption.
All clients can expect 24/7 enterprise-level support. Available through chat, phone, and we'll be in-person within 72 hours for any SEV1 issues.
No. We offer paid pilots for evaluation purposes.
You do. We believe all outputs, derivatives, and insights belong to the client. Our platform is to help you get there quicker.
We can get BioBox fully deployed within 7 days.



